Privacy-preserving Large language models for
Acronym Clinical Inference and Disambiguation

Privacy-preserving Large language models for
Acronym Clinical Inference and Disambiguation
Interactive Demo
Clinical note
The reports SOB after exertion. Prior noted in chart. Plan includes consult after discharge and repeat in the morning.
Selected acronym
PT is interpreted as patient in this sentence because the note says the PT "reports" symptoms.
Click an acronym to see how PLACID resolves it from context.
PLACID answer
PT → Patient
The phrase "reports SOB after exertion" makes PT most likely to mean patient, not physical therapy.
Context selector
"PT tolerated gait training and stairs."
The Problem
Electronic health records contain tens of thousands of acronyms. "PT" could mean patient, physical therapy, prothrombin time, or posterior tibial, depending on context. Misinterpretation contributes to clinical errors and hinders downstream NLP tasks.
Existing solutions require sending sensitive patient data to external APIs, a non-starter in healthcare. PLACID addresses both problems at once.
Acronym Ambiguity Example
Distribution depends entirely on clinical context
The Approach
Runs entirely on-premises. Patient data never leaves the institution. No external API calls, no data sharing.
Uses surrounding clinical context, the full sentence, section headers, patient history, to infer the most likely acronym expansion.
Designed to operate at scale across millions of clinical notes, improving downstream NLP tasks including coding, summarisation, and QA.
The Interface
A clinical note is pasted or uploaded, PLACID identifies and annotates every acronym inline, then returns a disambiguation summary with per-acronym confidence scores and domain classifications, all processed locally, no data leaves the institution.
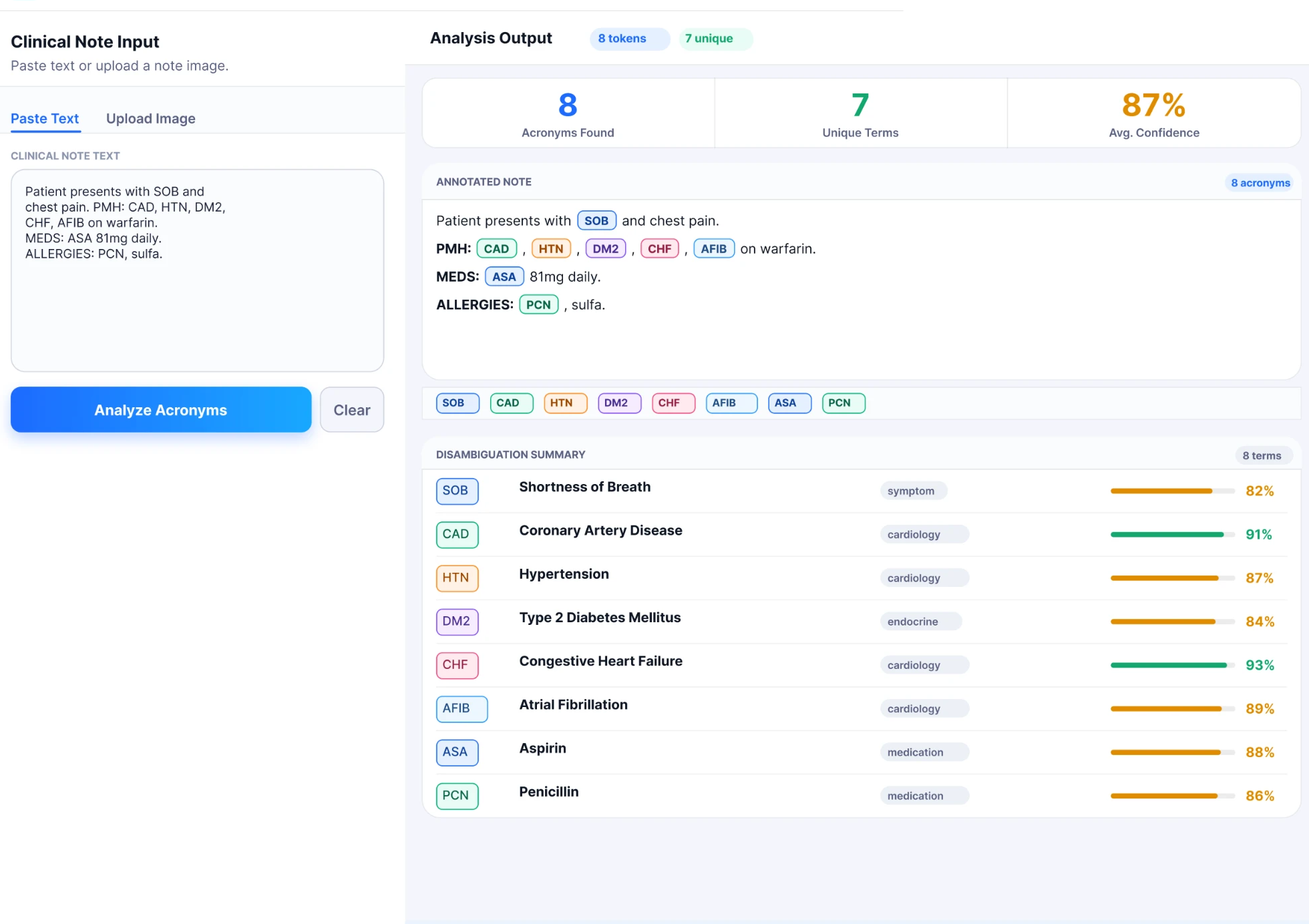
PLACID interface · Clinical note analysis with inline annotation and disambiguation summary · Figma prototype
PLACID is being integrated directly into Electronic Health Record workflows, running inline disambiguation on clinical notes at the point of documentation. By operating entirely on-device, it meets strict healthcare data governance requirements without requiring any external API calls or data transmission.
Clinical notes and discharge summaries shared with patients are dense with acronyms that are meaningless to non-clinicians. PLACID is being adapted for patient-facing portals and after-visit summaries, automatically expanding and contextualising clinical abbreviations to improve health literacy and reduce patient confusion.
Performance
Disambiguation Accuracy by Acronym Frequency
* Figures are representative, see arXiv paper for full experimental details.
Implementation Summary
| Overall accuracy | See arXiv paper for the complete evaluated breakdown; headline results are shown above by acronym frequency. |
|---|---|
| Model size range | Local LLM variants selected to balance accuracy, latency, and on-premises deployment constraints. |
| Deployment platform | Apple M4 Max silicon with MLX framework model optimization. |
| Privacy boundary | Clinical notes remain local; no external API transfer is required. |
0
Acronyms in scope
0
% Local inference
0ms
External data transfer
0
Team members
Team

Manjushree B. Aithal
Lead Author · PhD
Department of Biomedical Informatics · CU Anschutz

Alexander Kotz
Co-author · CPBS PhD Student
Computational Bioscience · CU Anschutz

James Mitchell
Senior Author · PI
Department of Biomedical Informatics · CU Anschutz
arXiv:2603.23678 · March 2026 · Accepted poster, AMIA Symposium 2026
Available on arXiv and accepted as a poster for the AMIA Symposium 2026.